RNA FISH for stem cell research
Stem cells hold great promise for the treatment of various diseases due to their capacity for self-renewal and ability to differentiate. Stellaris™ RNA fluorescence in situ hybridisation offers a unique insight into the expression patterns of embryonic and adult stem cells.
Visualisation, quantification, and localization of individual RNA molecules provide insight into the unique expression patterns of embryonic and adult stem cells. Though stem cells hold great promise for the treatment of various diseases due to their capacity for self-renewal and ability to differentiate, a basic understanding of stemness remains elusive. Advanced methods of analysis are required for the complex gene expression present in stem cells.
Custom or DesignReady Probe sets
LGC, Biosearch Technologies‘ free, online Stellaris Probe designer allows you to craft Stellaris RNA FISH probes with optimal binding properties for your target RNA sequence. A custom Stellaris RNA FISH probe set is a blend of up to 48 oligos labelled with a fluorophore.
We also offer DesignReady probe sets which are professionally designed and go through rigorous bioinformatics analysis to ensure specificity. These made-to-order probe sets include stem cell-specific targets across a wide range of model organisms. More information and current protocols are available online.
Example DesignReady Probe sets |
|
| BCL2 | GFAP |
| CDKN1A | NANOG |
| KIT | NES |
| NOTCH1 | POU5F1 |
| TERT | SOX2 |
| XIST | H19 |
| NEAT1 | HOTAIR |
Detect, localise, and quantify RNA molecules in cells or intact tissue
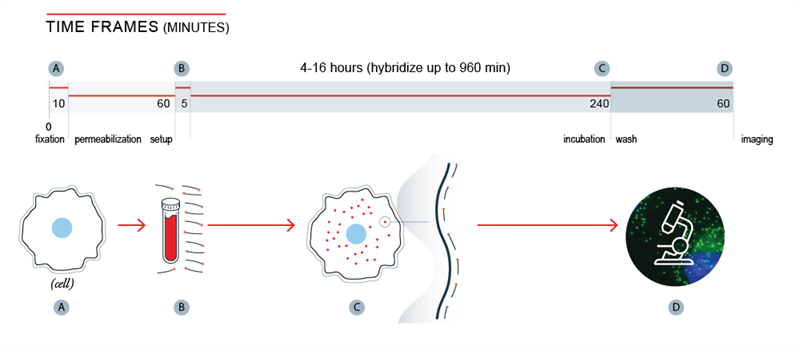
Biosearch Technologies’ Stellaris RNA FISH probes, can quantify and localise RNA within cells, tissue sections, or whole mount tissue where the morphology is preserved. A Stellaris FISH probe set comprises multiple oligonucleotides targeting a single RNA target. The fluorescently labeled probes bind along the target transcript, and combine to produce a punctate signal for an individual RNA molecule.
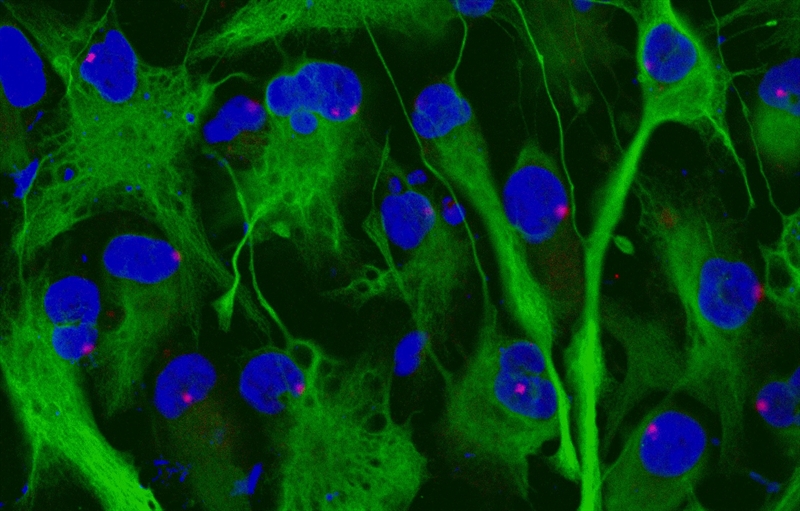
Image of Stellaris RNA FISH with Immunofluorescence. XIST (red), GFAP protein (green). Photo credit: N. Khoury
Publication highlights
Over 33% of publications citing Stellaris RNA FISH publish in Cell, Nature, or Science
Visit our Citation Center for the complete list.
Stem cell publication highlights
Wnt Ligands Secreted by Subepithelial Mesenchymal Cells Are Essential for the Survival of Intestinal Stem Cells and Gut Homeostasis
Valentaet al. Cell Reports. 2016. url: https://dx.doi.org/10.1016/j.celrep.2016.03.088
Dissecting Transcriptional Heterogeneity in Pluripotency: Single Cell Analysis of Mouse Embryonic Stem Cells
Guedeset al. Methods in Molecular Biology. 2016. url: https://dx.doi.org/10.1007/7651_2016_356
Transposon Dysregulation Modulates dWnt4 Signaling to Control Germline Stem Cell Differentiation in Drosophila
Upadhyayet al. PLOS Genetics. 2016. url: https://dx.doi.org/10.1371/journal.pgen.1005918
CD133 does not enrich for the stem cell activity in vivo in adult mouse prostates
Weiet al. Stem Cell Research. 2016. url: https://dx.doi.org/10.1016/j.scr.2016.03.003
High ImpactSingle-cell differences in matrix gene expression do not predict matrix deposition
Coteet al. Nature Communications. 2016. url: https://dx.doi.org/10.1038/ncomms10865
High ImpactReplacement of Lost Lgr5-Positive Stem Cells through Plasticity of Their Enterocyte-Lineage Daughters
Tettehet al. Cell Stem Cell. 2016. url: https://dx.doi.org/10.1016/j.stem.2016.01.001
X chromosome inactivation in human parthenogenetic embryonic stem cells following prolonged passaging
High ImpactProteostatic Control of Telomerase Function through TRiC-Mediated Folding of TCAB1
Freund et al. Cell. 2014. url: https://dx.doi.org/10.1016/j.cell.2014.10.059
High ImpactStochastic promoter activation affects Nanog expression variability in mouse embryonic stem cells
Ochiai et al. Nature Scientific Reports. 2014. url: https://dx.doi.org/10.1038/srep07125
High ImpactLuminal signalling links cell communication to tissue architecture during organogenesis
Durdu et al. Nature Letter. 2014. url: https://dx.doi.org/10.1038/nature13852
Defined Human Pluripotent Stem Cell Culture Enables Highly Efficient Neuroepithelium Derivation Without Small Molecule Inhibitors
Lippmann et al. Stem Cells. 2013. url: https://dx.doi.org/10.1002/stem.1622
A dynamic population of stromal cells contributes to the follicle stem cell niche in the Drosophila ovary
Identifying Division Symmetry of Mouse Embryonic Stem Cells: Negative Impact of DNA Methyltransferases on Symmetric Self-Renewal
Jasnos et al. Stem Cell Reports. 2013. url: https://dx.doi.org/10.1016/j.stemcr.2013.08.005
SNP-FISH, High ImpactAllele-specific detection of single mRNA molecules in situ
Hansen et al. Nature Communications. 2013. url: https://dx.doi.org/10.1038/nmeth.2601
linc-HOXA1 is a noncoding RNA that represses Hoxa1 transcription in cis
Maamar et al. Genes & Development. 2013. url: https://dx.doi.org/10.1101/gad.217018.113
Single-cell gene expression profiling reveals functional heterogeneity of undifferentiated human epidermal cells
Tan et al. Development and Stem Cells. 2013. url: https://dx.doi.org/10.1242/dev.087551
High ImpactAsymmetric Segregation of the Double-Stranded RNA Binding Protein Staufen2 during Mammalian Neural Stem Cell Divisions Promotes Lineage Progression
Kuseket al. Cell Stem Cell. 2012. url: https://dx.doi.org/10.1016/j.stem.2012.06.006
High ImpactTPP1 OB-Fold Domain Controls Telomere Maintenance by Recruiting Telomerase to Chromosome Ends
Zhonget al. Cell. 2012. url: https://dx.doi.org/10.1016/j.cell.2012.07.012
The P granule component PGL-1 promotes the localization and silencing activity of the PUF protein FBF-2 in germline stem cells
Voronina et al. Development and Stem Cells. 2012. url: https://dx.doi.org/10.1242/dev.083980
High ImpactEGFR/Ras/MAPK Signaling Mediates Adult Midgut Epithelial Homeostasis and Regeneration in Drosophila
Jiang et al. Cell Stem Cell. 2011. url: https://dx.doi.org/10.1016/j.stem.2010.11.026