Stellaris getting started guide
This handbook provides tips and guidelines to help you appropriately plan and seamlessly perform Stellaris experiments.
learn moreThis handbook provides tips and guidelines to help you appropriately plan and seamlessly perform Stellaris experiments.
learn moreThese protocols are developed by the Stellaris™ Team to help you perform RNA FISH on various sample types.
learn moreView Stellaris images contributed by customers and by scientists from LGC, Biosearch Technologies.
learn moreFind peer reviewed publications citing Stellaris™ RNA FISH probe sets sorted by product, application, model organism and more.
learn moreFind answers to questions about how Stellaris works, the sensitivity and specificity, multiplexing, how to order and more.
learn moreDesign your own custom Stellaris™ RNA FISH Probes using our online Stellaris Probe designer, free with registration. By using this probe designer, you are just a few clicks away from placing your Stellaris order online.
learn moreStellaris™ RNA FISH Probes are used for single molecule RNA detection by fluorescence in situ hybridisation (FISH). View our custom design options, in house-designed probe sets, controls and buffers.
learn moreExplore the unique applications of Stellaris™ RNA FISH.
learn moreScientist-generated technical articles on diverse RNA FISH topics including probe set design, microscope use and imaging, proper controls, and special topics such as vision research, fusion-FISH, lncRNA localization and many others.
learn moreFind details about all the dyes and modifications currently offered for Stellaris RNA FISH Probe sets. Multiplexing recommendations, dye stability testing, emission and absorption spectra, use notes, publication links, and much more are here to help you choose the optimal modifications for your experiment.
learn moreHere, you will find answers to frequently asked questions about the Stellaris RNA FISH technology.
learn moreFind videos on topics including general Stellaris RNA FISH information, Multiplexing with IF, and webinars on Detection of lncRNA and advances in RNA characterisation.
learn more
Stellaris RNA FISH (fluorescence in situ hybridisation) is a RNA visualisation method that allows for simultaneous detection, localization, and quantification of individual RNA molecules at the cellular level in fixed samples using fluorescence microscopy. A set of Stellaris RNA FISH Probes is comprised of a pool of up to 48 unique probes, each labeled with a fluorophore, that collectively bind along an RNA target transcript to produce a diffraction limited spot.
The Stellaris RNA FISH technology follows a simple protocol, does not need exotic reagents and is platform-independent. It can offer same-day results and is versatile in terms of sample type, species and application. Stellaris RNA FISH probes can be labeled with several different dyes to allow for simultaneous detection of different RNA targets in multiplexed experiments. The Stellaris RNA FISH technology enables scientists to visualise gene expression through direct detection without isolation, purification or amplification.
The Stellaris RNA FISH protocol is simple, consisting of four main steps, as shown in the image below. For some sample types, the entire process can be completed in less than a day.
Step A: Prepare sample
A sample is grown, adhered, or sectioned onto a #1 cover glass and permeabilised with alcohol. No protease required!
Note: Variations in sample preparation between different organisms and sample types are covered in our online Stellaris protocols.
Step B: Hybridize probes
Hybridization can be completed in 4 to 16 hours at +37 °C in a standard laboratory incubator.
Step C: Wash sample
Removal of Hybridization Buffer and addition of wash buffers with short incubation periods are able to remove excess probes. The total time for this step is 1-1.5 hours.
Step D: Image sample
The sample is ready for imaging using a standard fluorescence microscope. We recommend using a wide-field fluorescence microscope though other imaging platforms can be used with Stellaris RNA FISH probes. Image processing is done using available online tools which convert Z stack images to a maximum intensity projection image.
The large number of probes in a Stellaris RNA FISH assay ensures both high sensitivity and specificity. The inherent redundancies of this direct detection method minimize the possibility of false negative and false positive signals. Positive signals are identified from the combined localized fluorescence of multiple probes. Off-target binding of single probes generates only weak and diffuse fluorescence, which falls well below the threshold for detection of the targeted mRNAs. The short probe lengths are able to easily penetrate the cellular matrix for efficient hybridization.
Stellaris RNA FISH probes are available in a wide variety of fluorophores in the visible spectrum, including LGC, Biosearch Technologies’ CAL Fluor™ and Quasar™ dyes and can be used in a multiplex assay. The dye choices in a multiplex assay are largely determined by the filters available on the imaging platform, but it is certainly possible to design an assay including a DAPI nuclear stain and three Stellaris RNA FISH probe sets labeled with reporter dyes with minimal spectral overlap. See our Stellaris Dyes and Modifications page for detailed information on all our available dyes and modifications as well as a multiplexing guide.
Multiplex assays can be used to identify a correlation in expression levels between multiple genes, compare a control probe set of known expression to the expression of a target, or simply save time over running the many separate assays. Our ShipReady Probe Sets are assays designed to detect well-known, previously validated targets and have been wet bench tested for success as positive controls. The ShipReady probe sets are convenient and affordable. Our DesignReady Probe Sets are made to order and take the guess work out of designing your own Stellaris assay. We have a catalogue of over 4,000 professionally designed assays for many common species and model organisms. These have undergone an additional level of bioinformatics analysis to ensure specificity.
Stellaris RNA FISH can also be combined with existing technologies such as qPCR, DNA FISH, IHC (immunohistochemistry), IF (immunofluorescence) and western blotting to provide complementary information.
Design your own Custom Probe Sets using our online Stellaris Probe Designer, free with registration. Follow the guidelines for sequence requirements on one of our Stellaris Tech Blogs: Designing Stellaris RNA FISH Probes Part I: Increasing Probe Count.
Finally, don’t forget to order a set of our Stellaris Buffers which enhance signal and reduce background.
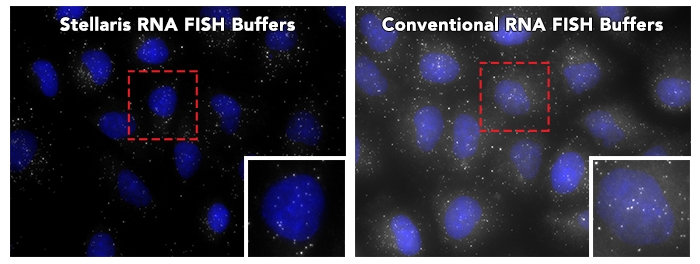
Stellaris RNA FISH Probes are sold under license from Rutgers, The State University of New Jersey, and may be used under its patent rights for Research Use Only.